Antibodies against double stranded DNA (anti-dsDNA) are immunological indicators for systemic lupus erythematosus (SLE), a severe rheumatic autoimmune disease. According to the recommendations of the American College of Rheumatology, their detection constitutes an important criterion for the diagnosis of the disease [1].
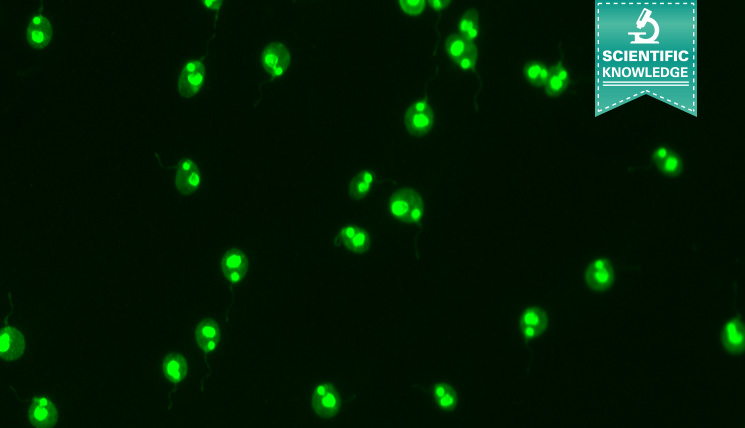
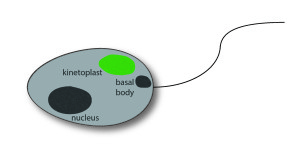 Among the commonly applied test systems, indirect immunofluorescence on the flagellate Crithidia luciliae is considered as particularly specific method to detect anti-dsDNA in patient samples. A positive reaction manifests in a fluorescent kinetoplast, a network of tightly packed dsDNA within the mitochondrion of the protist which is free of nuclear proteins, and therewith well suited for the selective detection of anti-dsDNA.
Among the commonly applied test systems, indirect immunofluorescence on the flagellate Crithidia luciliae is considered as particularly specific method to detect anti-dsDNA in patient samples. A positive reaction manifests in a fluorescent kinetoplast, a network of tightly packed dsDNA within the mitochondrion of the protist which is free of nuclear proteins, and therewith well suited for the selective detection of anti-dsDNA.
 Meanwhile, new automation platforms help to perform IIFTs efficiently and objectively: The EUROPattern Suite is an elaborate system, encompassing an automated microscope and specific classification software, which has already proven its value in the fully automated evaluation of fluorescence patterns of anti-nuclear antibodies (ANA) on HEp-2 cells [2, 3]. Now, the EUROPattern Suite can also evaluate Crithidia luciliae (anti-dsDNA) immunofluorescence tests (CLIFT). The corresponding CLIFT classification software was validated on the basis of 669 samples (569 consecutive samples, sent to the Clinical Immunological Laboratory, Lübeck, with request for anti-dsDNA testing and 100 samples of healthy blood donors) in the recently published study by Gerlach et al. [4]. The automated software evaluation of the CLIFT was compared to its visual evaluation.
Meanwhile, new automation platforms help to perform IIFTs efficiently and objectively: The EUROPattern Suite is an elaborate system, encompassing an automated microscope and specific classification software, which has already proven its value in the fully automated evaluation of fluorescence patterns of anti-nuclear antibodies (ANA) on HEp-2 cells [2, 3]. Now, the EUROPattern Suite can also evaluate Crithidia luciliae (anti-dsDNA) immunofluorescence tests (CLIFT). The corresponding CLIFT classification software was validated on the basis of 669 samples (569 consecutive samples, sent to the Clinical Immunological Laboratory, Lübeck, with request for anti-dsDNA testing and 100 samples of healthy blood donors) in the recently published study by Gerlach et al. [4]. The automated software evaluation of the CLIFT was compared to its visual evaluation.
In the initial step, the samples were incubated on the BIOCHIPs of the commercial Crithidia luciliae (anti-dsDNA) IIFT. Subsequently, the EUROPattern microscope took focused images of the incubated slides (one image per BIOCHIP), automatically. These were then evaluated both visually (by two independent experts; in case of disagreement, evaluation of a third expert was decisive) and automatically by the EUROPattern Suite. The cells and BIOCHIPs, respectively, were classified as anti-dsDNA positive or negative according to the fluorescence status of the kinetoplast. A comparison of the software-generated and the visual classification revealed that the EUROPattern Suite assessed 577 of 596 (visually) negative samples as likewise negative (specificity 96.8%). Automated classification of positive samples (n = 73) was 100% accordant to visual classification (sensitivity 100%). Therewith, the laboratory physician may trust in the system to detect positive samples reliably and to not issue false negative findings.
| n = 669 | Visual evaluation | |||
| positive | negative | Σ | ||
| EUROPattern evaluation | positive | 73 | 19 | 92 |
| negative | 0 | 577 | 577 | |
| Σ | 73 | 596 | 669 | |
Overview of the functionality of the system
The software merges single results from different sample dilutions into one final report (classification, titer estimation) per patient. This report is given on the computer screen together with a confidence value and the individual images. Here, the laboratory expert has to check and confirm the result. For the benefit of efficiency, all negative results may be authorised by few mouse clicks. The utilised graphical user interface is a component of the superordinate laboratory management software EUROLabOffice (ELO). ELO archives all digitally confirmed results and generates a comprehensive patient history which also incorporates results from further analyses. For this purpose, the software organises the complete data exchange between the different laboratory workstations (IIFT, immunoblot, ELISA …) and the standard laboratory information systems.
Thus, the EUROPattern Suite is able to support the laboratory personnel in the evaluation of IIFT and to considerably reduce the manual work load. At the same time, subjectivity in the interpretation of the results is minimized by the automated classification while high quality is maintained compared to visual evaluation.
[1] Tan EM et al., Arthritis & Rheumatism 1982, 25(11): 1271–1277. [2] Voigt J et al., Clin Dev Immunol 2012, ID 651058. [3] Krause C et al., Lupus 2015, 24(4-5): 516–529. [4] Gerlach et al. J Immunol Res 2015, ID 742402.